 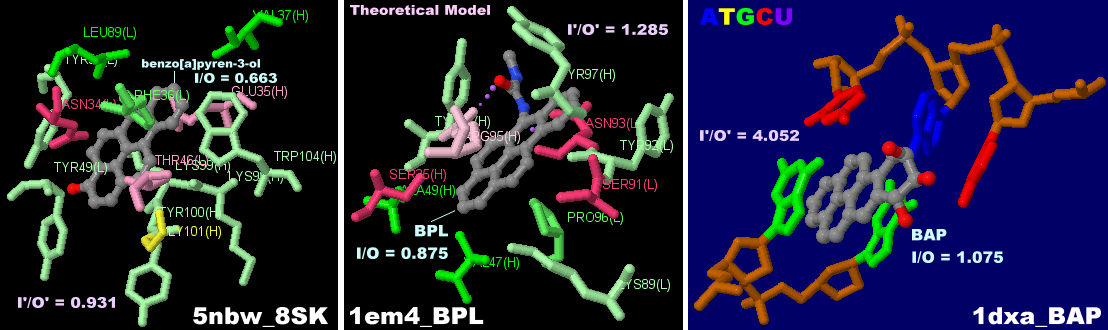
We have developed a computational approach to predict the structure of protein/ligand complexes based solely on the unbound conformation, the ligand, and easy-to-assess experimental data. In many cases, a structure of the unbound receptor is available, but conformational changes with respect to the ligand-bound form preclude it from being used as a basis for structure-based drug design. A critical prerequisite is a structure of the target protein close to its ligand bound conformation which is often difficult to determine experimentally. Structure-based drug design has become a powerful tool in modern drug discovery pipelines. A protocol is presented that is expected to enable structure modeling of protein/ligand complexes and structure-based drug design for cases where crystal structures of ligand-bound conformations are not available. In all cases, receptor models within 1.6 Å backbone RMSD to the target were predicted and close-to-native ligand binding poses were obtained for 8 of 10 cases in the top-ranked complex models. The method is applied to ten cases in which proteins undergo structural rearrangements of up to 7.1 Å backbone RMSD upon ligand binding. We present a method to predict the structure of protein/ligand complexes based solely on the apo structure, the ligand and the radius of gyration of the holo structure. Hence, if only an unbound (apo) structure is available distinct from the ligand-bound conformation, structure-based drug design is severely limited. Accurate receptor models in the ligand-bound state (holo structures), however, are a prerequisite for successful structure-based drug design.  
Computational methods for structure prediction of protein/ligand complexes, particularly docking, are as yet restricted by their limited consideration of receptor flexibility, rendering them not applicable for predicting protein/ligand complexes if large conformational changes of the receptor upon ligand binding are involved. Experimental structure determination of such complexes at atomic resolution, however, can be time-consuming and costly. Biological function of proteins is frequently associated with the formation of complexes with small-molecule ligands. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
